Dogs that are more related are likely to be more similar genetically than less related dogs. Breeders often want to know how related a dam and sire are because it tells them something about the traits likely to be expressed in a litter of puppies, and it also reflects the risk of inherited genetic disorders.
We can put a number on the degree of genetic similarity between a pair of dogs using the "Relationship Coefficient," which is the fraction of their genes that they share in common.
For example, a parent and offspring should have a relationship coefficient of 0.5, because the progeny inherits half of its genes from each parent. A pair of dogs that are less related, like half-sibs or cousins, will be less similar genetically and have a lower relationship coefficient.
You can estimate the relationship coefficient between a pair of dogs from pedigree data. Of course, you can never really know exactly which alleles have been inherited by a dog, so the relationship coefficient determined from a pedigree is just an estimate. You can get a far better estimate of genetic similarity by directly comparing the genes in the pair of dogs.
Thanks to DNA analysis, we can now do this.
ICB has been working on the development of new tools that will provide breeders with information about the genetic similarity of dogs by direct comparison of their DNA sequences. The resulting number for a pair of individuals is called the "genomic relationship coefficient". The higher this number, the more similar the DNA of the two dogs. (Identical twins would have a genomic relationship coefficient of 1.)
You might also be interested in the relatedness among the dogs in a group, or between two groups. Perhaps you want to compare dogs from two different subpopulations (e.g. bench vs field lines, UK vs US dogs, or dogs from different kennels). For this, you would determine the genomic relationship coefficients for every pair of dogs in the groups to be compared. If you only have a few dogs, this is relatively easy, but for more dogs the table of numbers gets big very quickly.
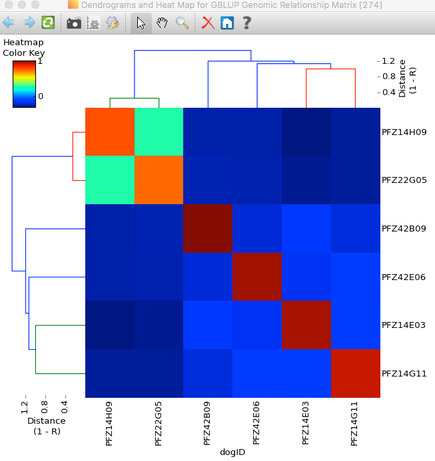
To make it easy visualize patterns in the relationship coefficients among larger numbers of dogs, we can use two graphical tools. The first is called a dendrogram, which is a type of pedigree tree that visually displays the magnitude of the relationship coefficient by the length of the branches. The other tool is called a "heat map", because it displays numerical values using different colors or variations in the intensity of color. Let's look at an example.
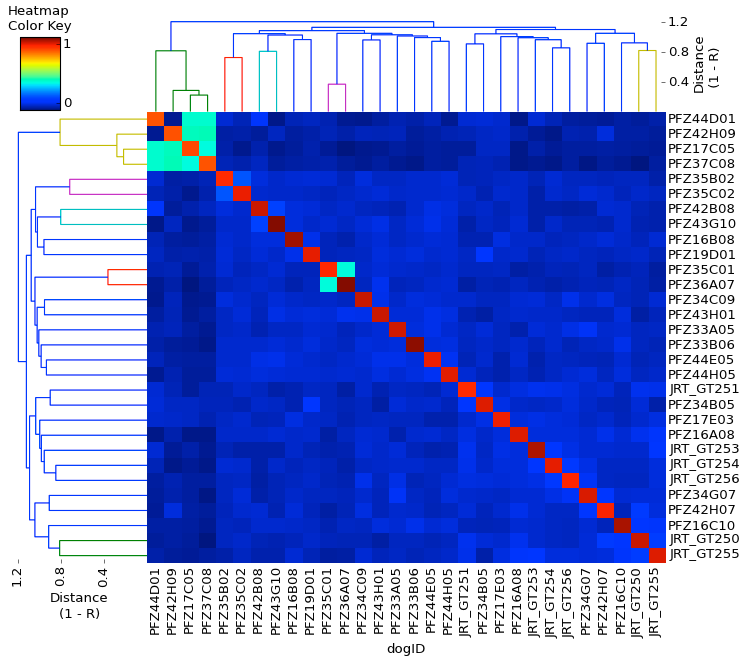
The IDs of the dogs are listed across the bottom and also down the right axis in the same order. The "Heatmap Color Key" in the upper left corner displays the colors that correspond to relationship coefficients from 0 to 1.
The row of red squares across the diagonal of the chart is each dog's relationship coefficient with itself, which is equal to one. The other squares are for each pairwise comparison of all of the dogs in the population. So, in the upper right hand corner, the first square compares dog PFZ44D01 with dog JRT_GT255. The square is a dark shade of blue, indicating that their genetic similarity is relatively low. In the middle of the graph you can see two turquoise squares, which indicate a pair of dogs that have a higher amount of genetic similarity. The reason there is a pair of squares on each side of the line of red squares is because the graph is a mirror image of itself along the diagonal. (Convince yourself of this by finding the IDs of the the pair of dogs for each of the turquoise squares.)
This chart provides you with a lot of information about this particular population of Jack Russel Terriers. There is a group of 4 dogs that seem to be genetically distinct from all of the rest, three of which are very closely related. The rest of the dogs vary in relatedness in pairwise comparisons, but on average their genetic similarity is low.
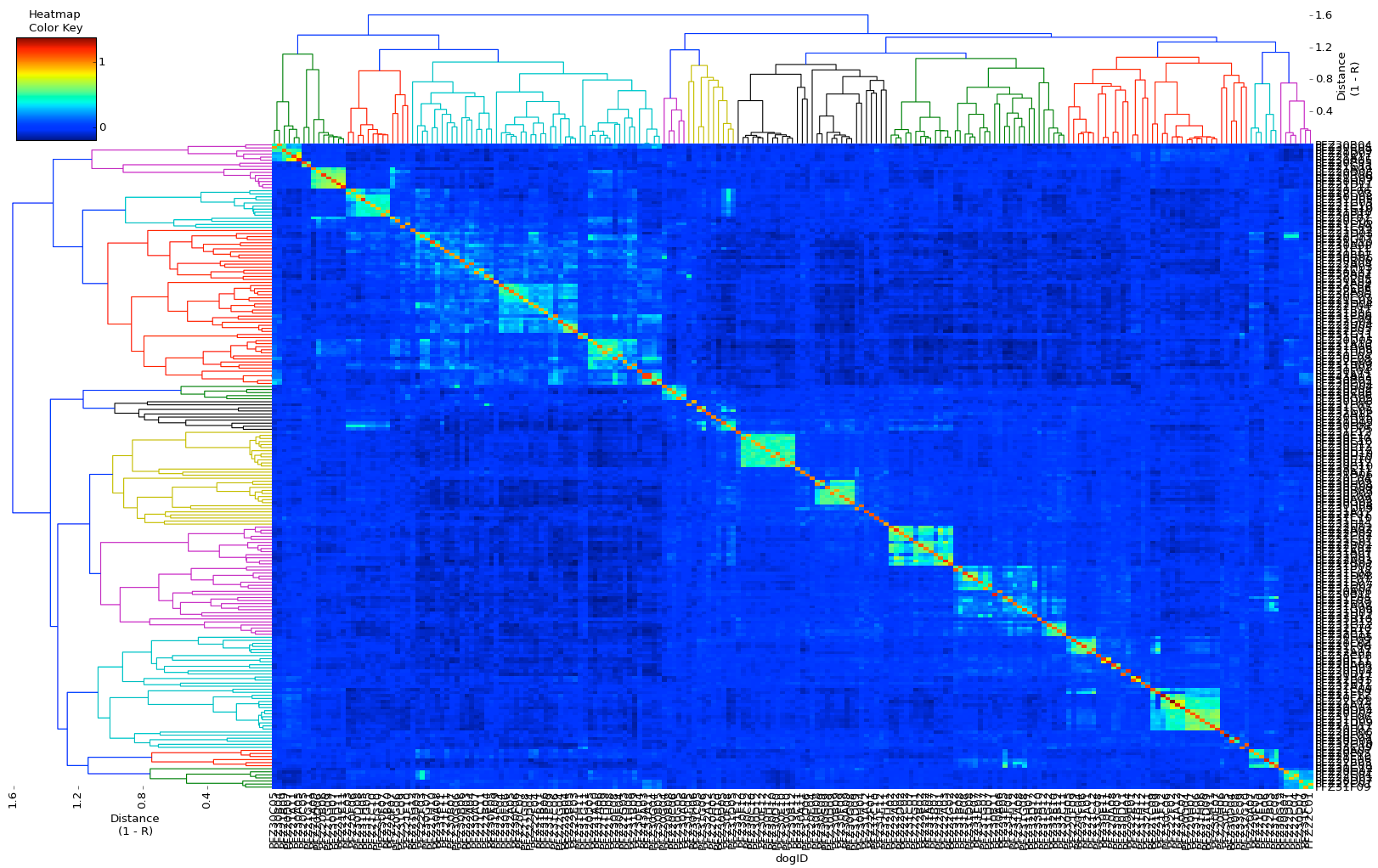
You should see in the Wolfhound data that there are actually two large groups of dogs connected by the highest horizontal branch. I don't have any information about these dogs, but the two clusters might depict dogs from different countries, or perhaps lines that have been bred more or less independently for many generations. The shortest branches might be connecting the dogs in a litter. The overall lightness of the blue (compared, for example, to the graph for Jack Russell Terriers) indicates that the genetic diversity in this population of dogs is relatively low.
Introducing: ICB's Genomic Breeding Tool
ICB's Genomic Breeding Tool is the most powerful genetic tool available to breeders today, providing information about:
- genetic similarity of pairs of dogs by direct comparison of DNA
- individual inbreeding coefficient of each dog
- overall genetic similarity of the dogs in the population
- identification of subpopulations or genetically similar clusters
- genetic diversity of the population
The DNA analysis ICB uses for this compares uses the newest version of the high density Illumina CanineHD SNP chip. This analysis identifies more than 200,000 DNA nucleotides distributed across each of the dog's 38 autosomal chromosomes, the two sex chromosomes, and the mitochondrial DNA (mtDNA). The same DNA analysis also tests for most genetic mutations for which a test is available, as well as X and Y chromosome and DLA haplotypes. This is the highest level of genetic comparison available anywhere. (For comparision, MyDogDNA uses about 7,000 nucleotides for their breeding tool, and the UC Davis VGL genetic diversity test uses 33 microsatellites.)
The data produced for the ICB Genomic Breeding Tool can also be combined with pedigree data, which will allow breeders to infer the relationships among dogs for which DNA is not available. We can also superimpose health information on this, which will facilitate identifying lineages of dogs at greater risk of a particular disorder. (Read about how to do this here.)
If you think your breed might be interested in participating in beta testing, please contact ICB.
ICB's online courses
*******************
Join our Facebook Group
ICB Breeding for the Future
...the science of dog breeding
*******************
Visit our Facebook Page
ICB Institute of Canine Biology
...the latest canine news and research