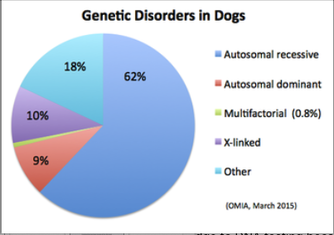
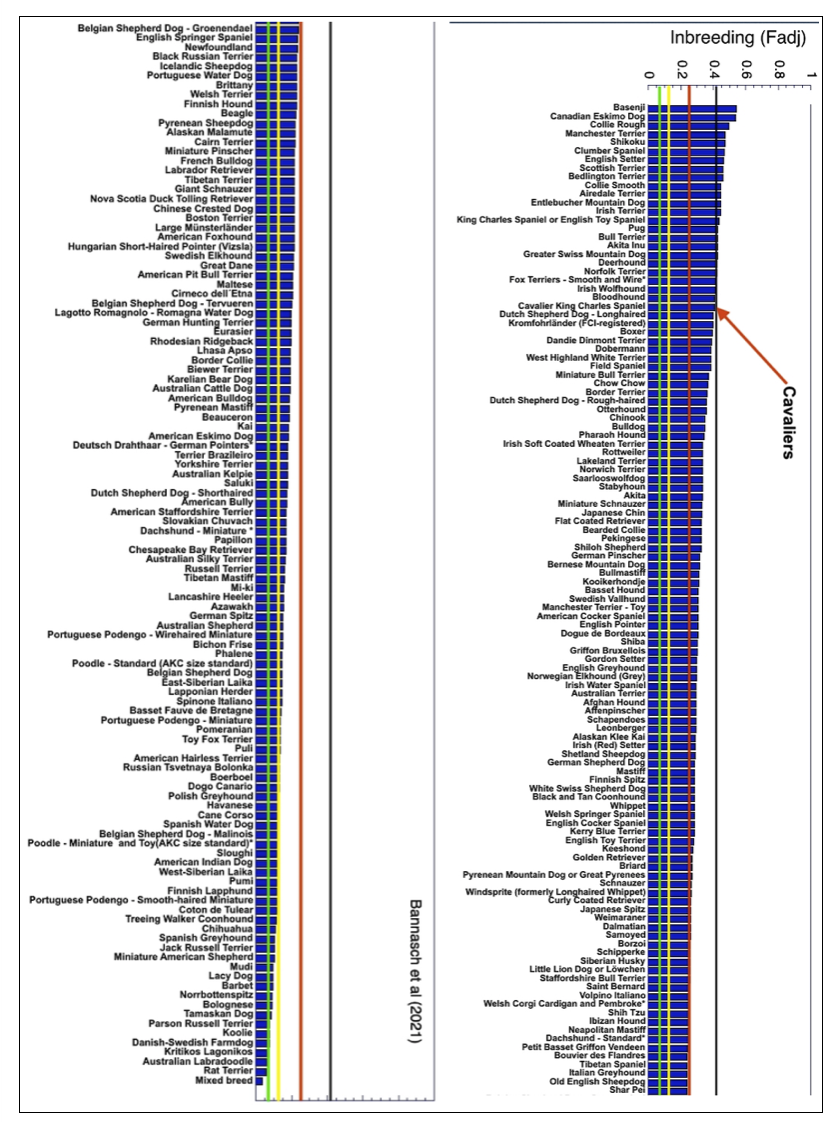
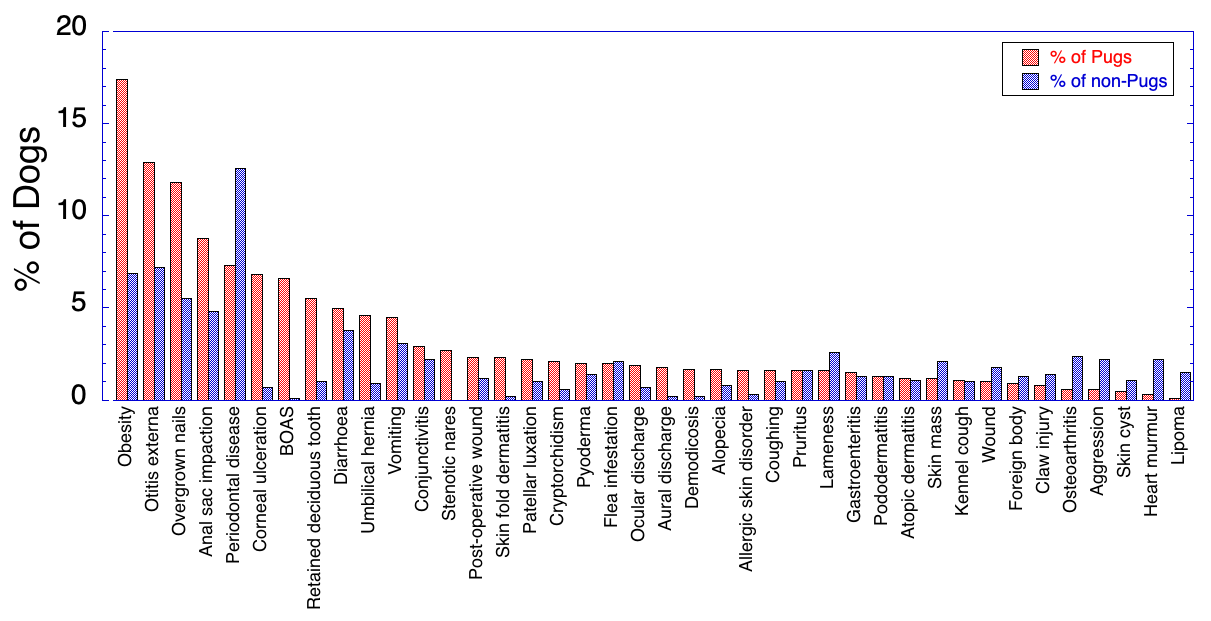
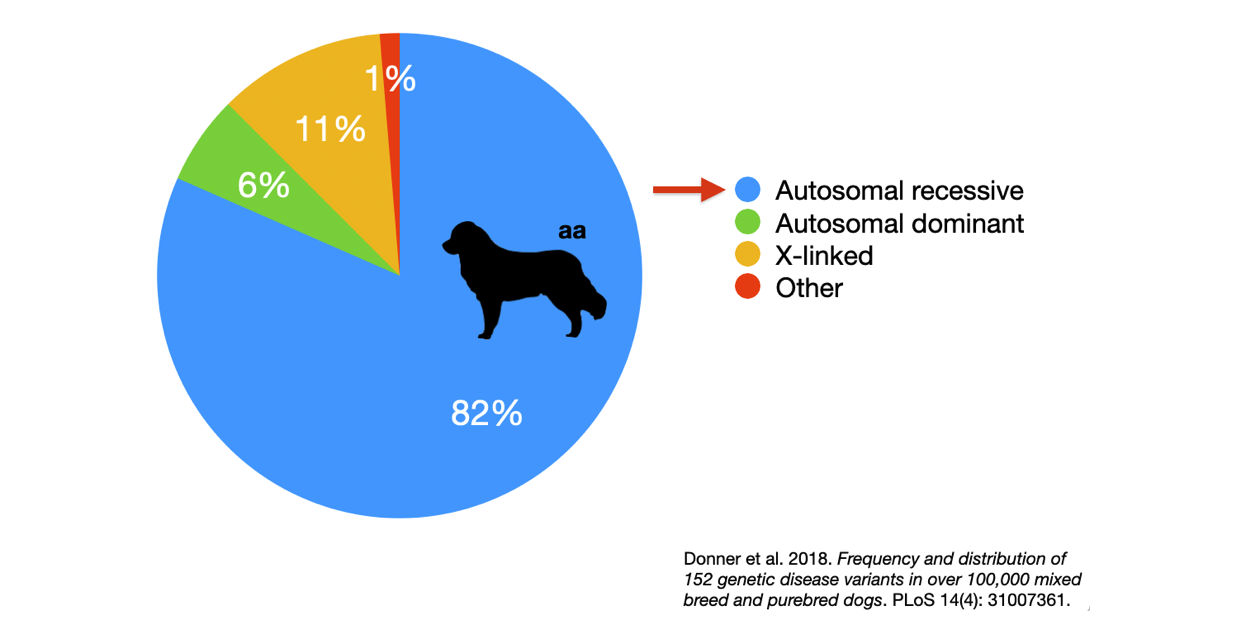
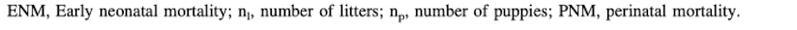6/16/2024 BREEDER MYTH: Crossbreeding will produce the disorders of both breeds in the puppies
5/9/2024 It's Westminster time again! But still, there's an elephant in the room
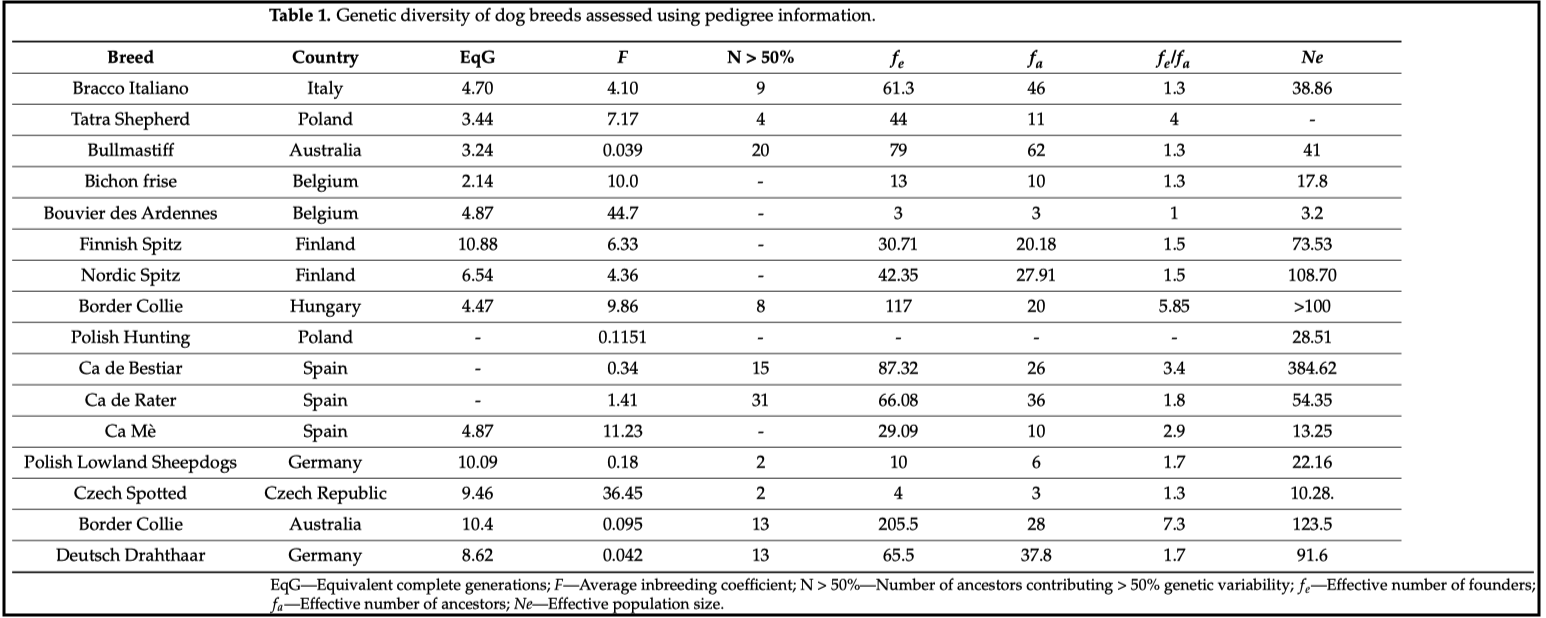
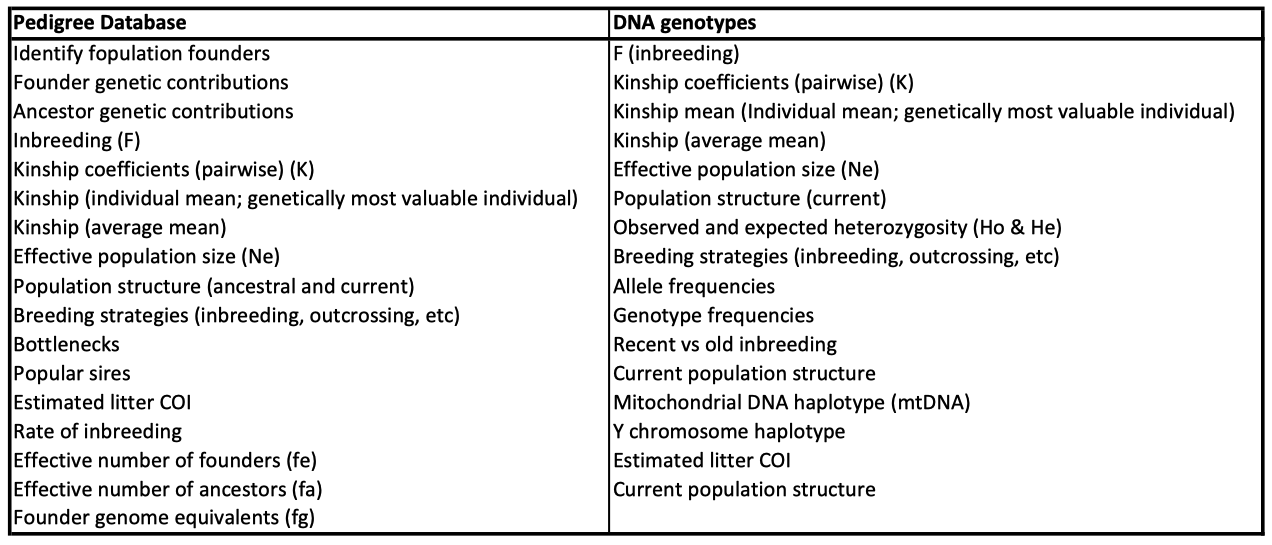
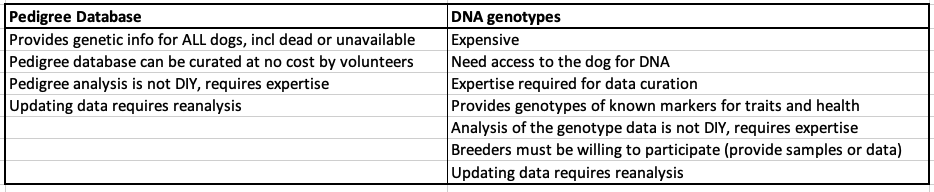
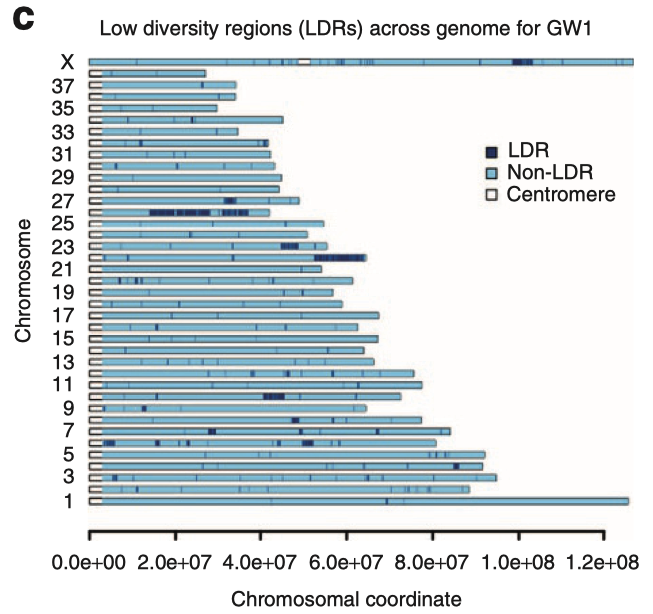
3/29/2024 Genetic management of purebred dogs: the tools you need in your toolbox
3/17/2024 Does your breed need genetic rehabilitation? Here's where to start
2/6/2023 What level of inbreeding is safe?
7/18/2023 Treating heat stress in dogs based on science
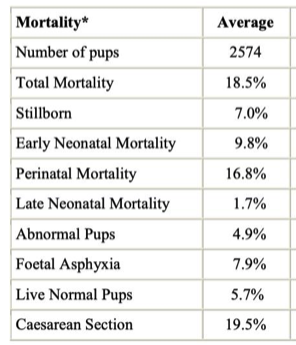
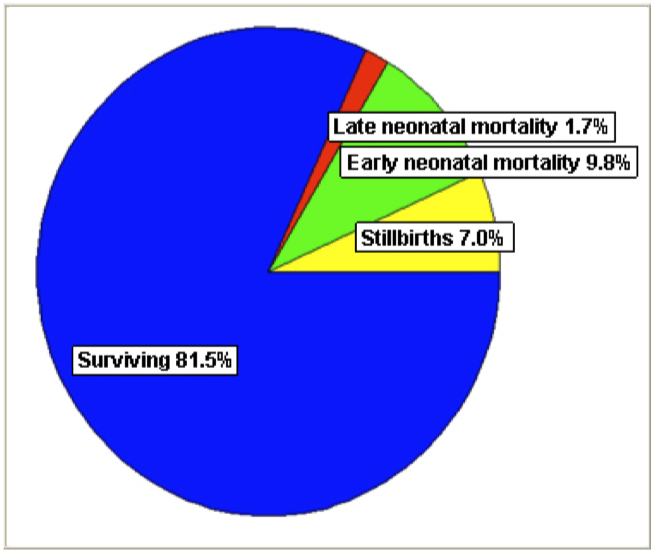
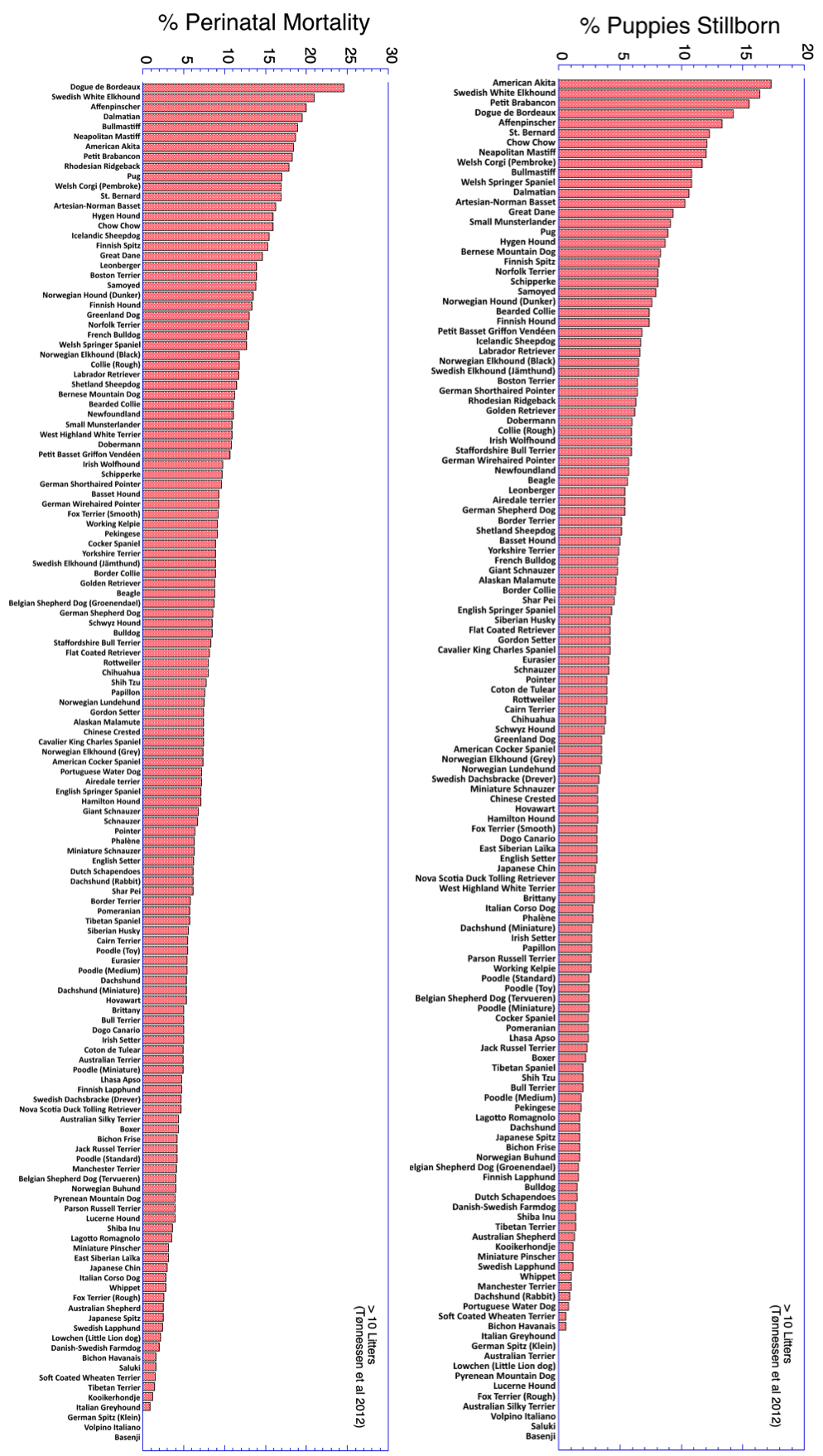
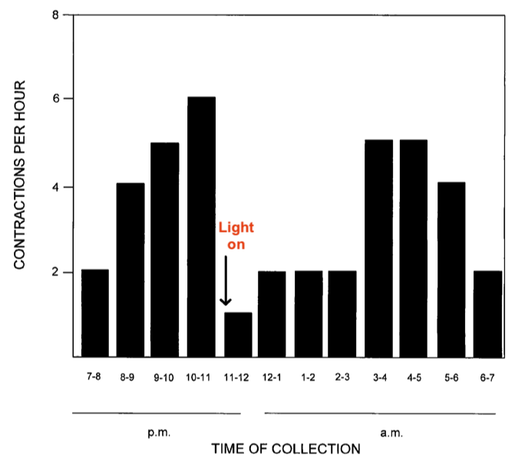
4/22/2023 We can prevent neonatal puppy mortality
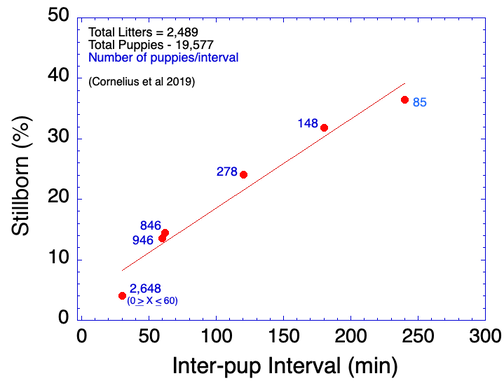
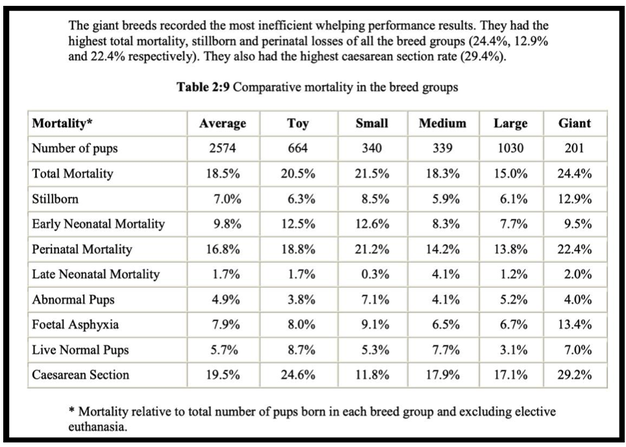
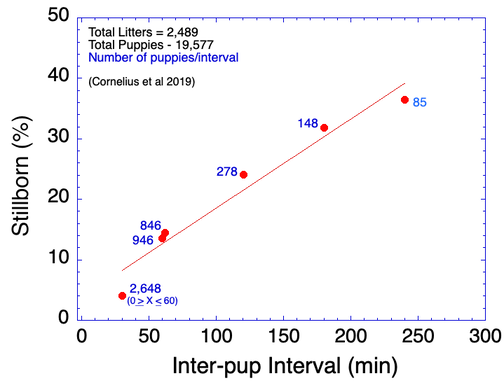
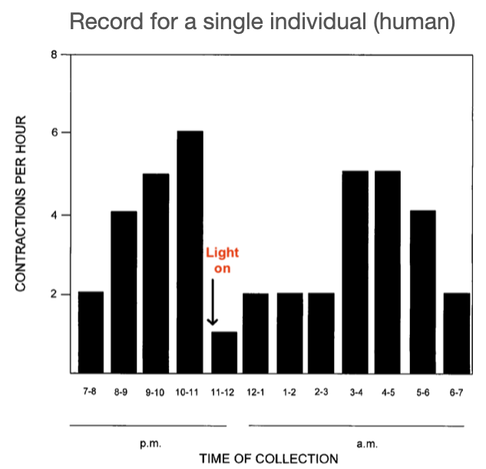
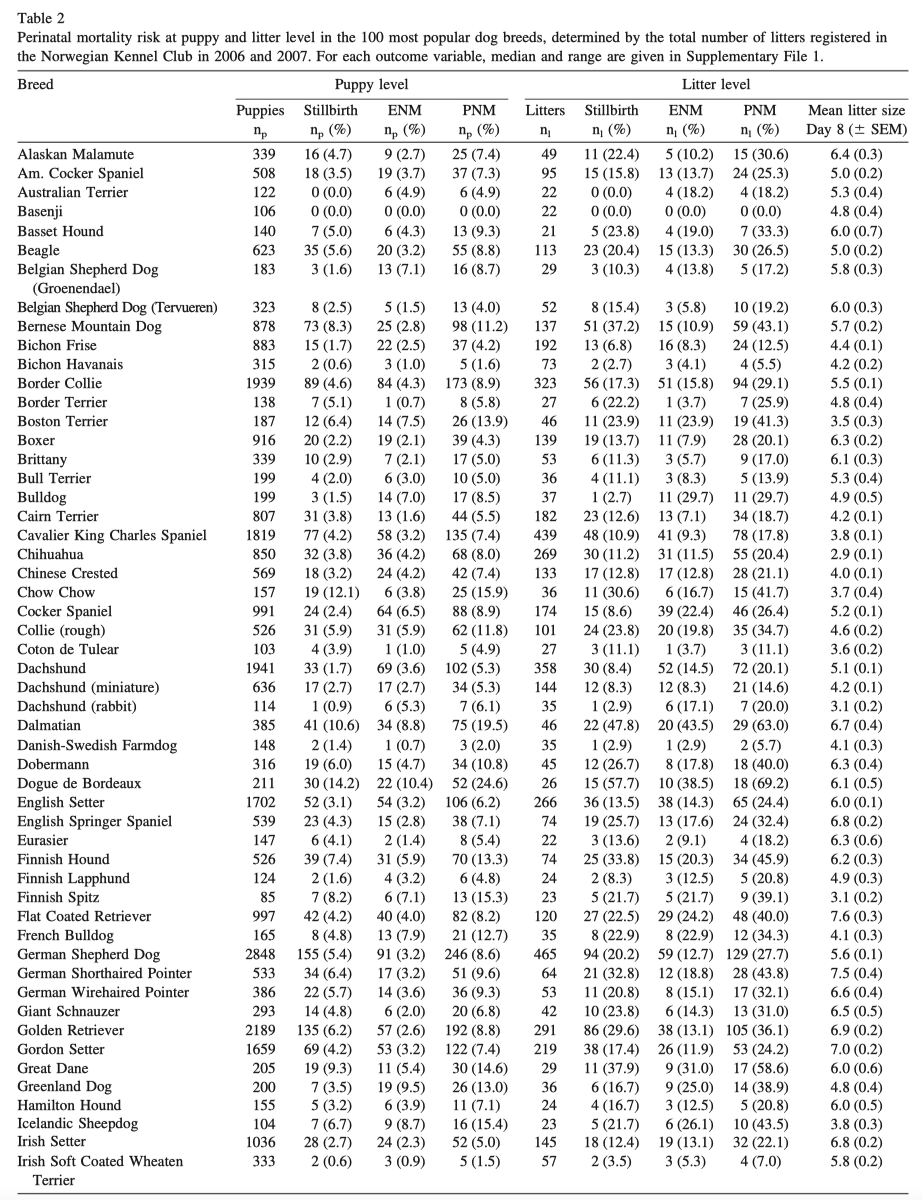
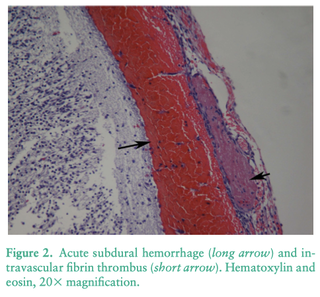
4/14/2023 Why do so many puppies die?
11/13/2022 Please don't swing the puppy
1/5/2022 The behavior of hot and cold puppies
7/31/2022 The "nonsense" of inbreeding coefficients and breeding restrictions on sires
5/22/2022 Is COI an essential tool or just a fad?
5/20,2022 The elephant in the room? Incest breeding.
5/19/2022 Is the Pug a "typical" dog?
4/10/2022 Breeding is managed evolution
3/22/2022 Puppies from a breed cross will have the health problems of both breeds: T or F?
3/20/2022 Do you know what you need to save your breed?
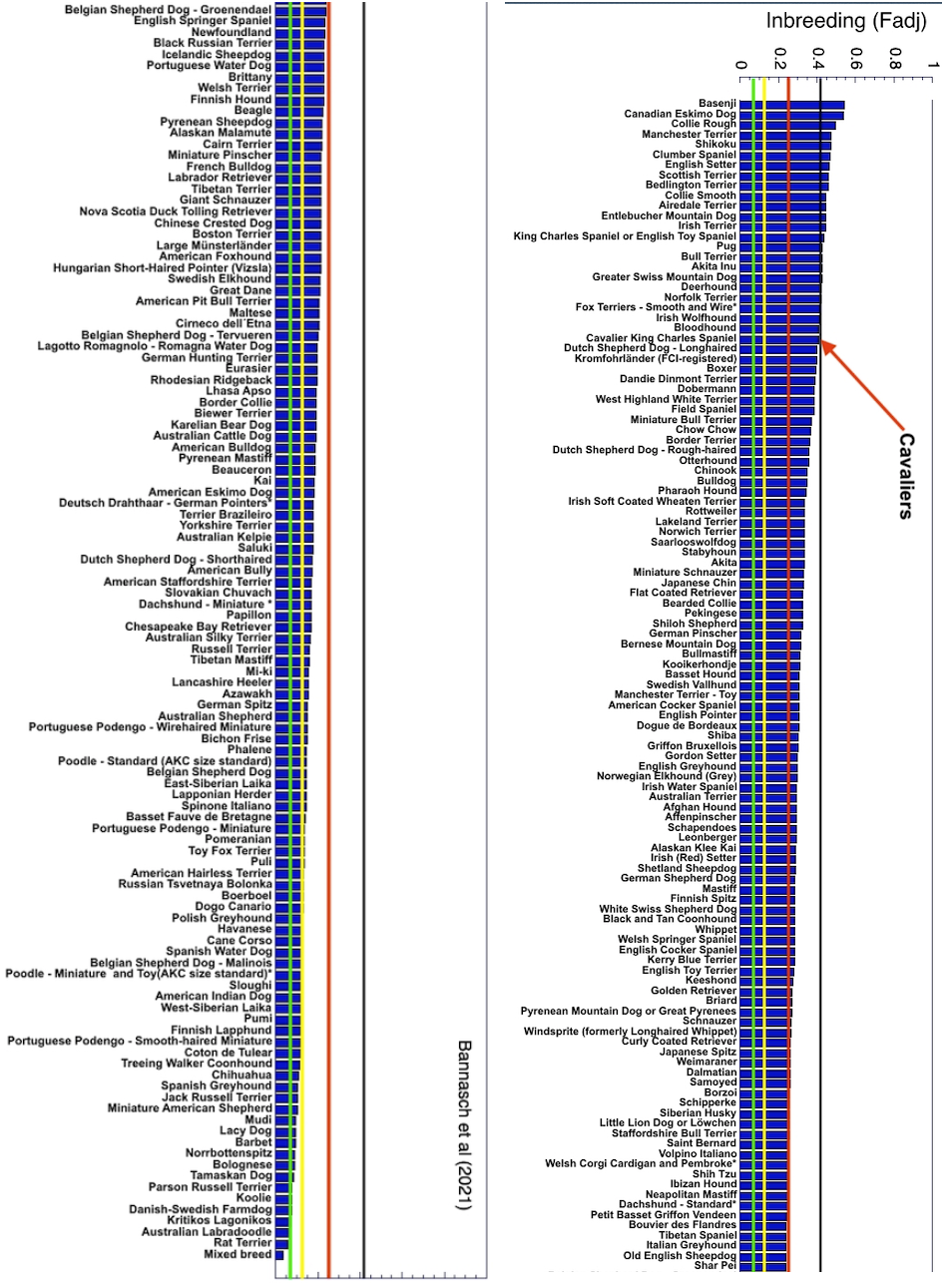
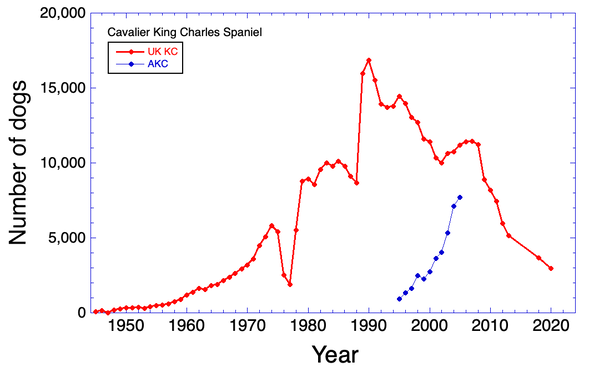
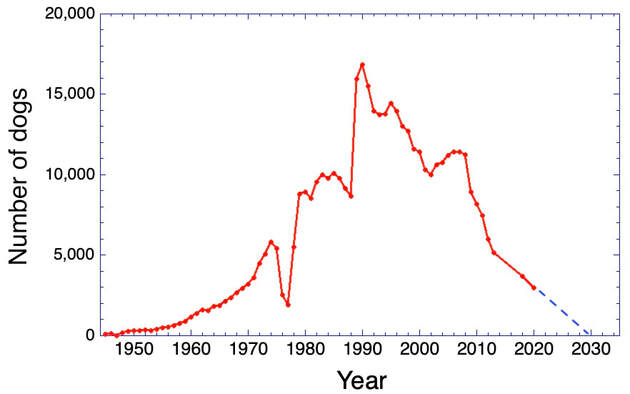
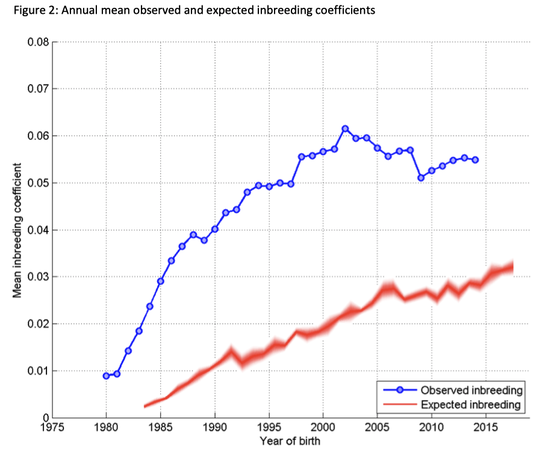
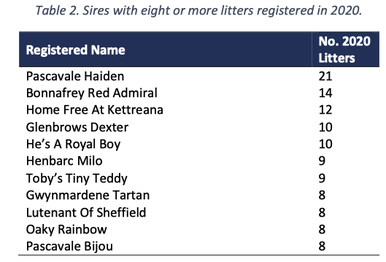
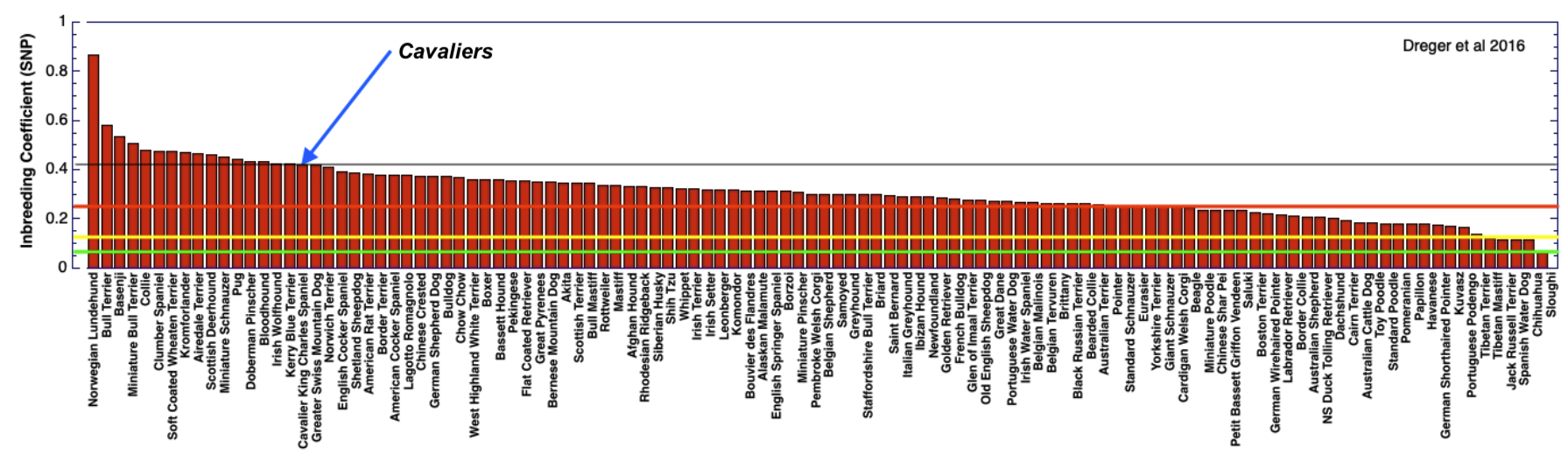
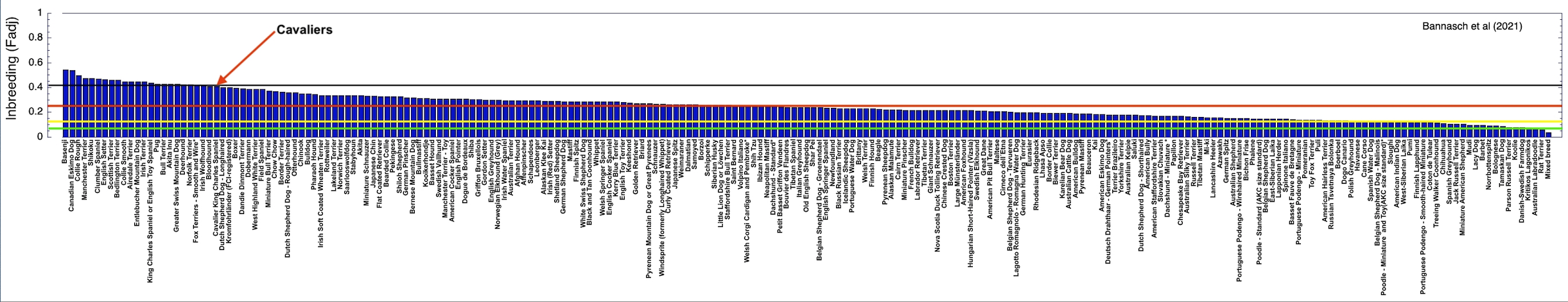
3/4/2022 The genetic status of the Cavalier King Charles Spaniel, part 1: Inbreeding
2/22/2022 The "catastrophe" of cross-breeding: meet the Borgis
2/20/2022 We need a Mayo Clinic for dogs
2/10/2022 Cavaliers are in trouble
2/5/2022 Hello breeders, this is Science
2/3/2022 The Norway breeding ban: what does the law say?
2/1/2022 Norway bans the breeding of Bulldogs and Cavaliers. Now what?
11/27/2021 Finally...a summary of canine coat color genetics
10/3/2021 The easy way to improve hips and elbows
12/29/2020 How hips form and Wolff's Law
12/27/2020 The basics of hip dysplasia in dogs
1/3/2020 Do your puppies have enough traction in the whelping box÷
8/23/2019 Why do mixed breed dogs have so many mutations?
8/15/2019 About pithy statements vs knowledge
7/9/2019 Let's kill the breeder myths!
7/5/2019 What is "heritability" and why do you need to know?
6/17/2019 Facts vs fear mongering
4/29/2019 Is health problem X in my breed caused by inbreeding and/or loss of genetic diversity?
4/19/2019 No, we have NOT found the mutation that causes breathing problems in brachycephalic dogs
4/21/2019 An update on hip dysplasia in dogs
4/20/2019 Addison's Disease and those doggone DLAs
3/26/2019 Genetic rescue and rehabilitation: I. Restoring genetic diversity of a breed
3/12/2019 The key requirement for preservation breeding
2/28/2019 The genetics of canine behavior goes molecular
2/14/2019 The messy science of assessing working ability in dogs
2/5/2019 A new ICB course that will use the DNA data from YOUR dog!
1/20/2019 How to breed dogs that are better than their parents: the genetics of continuous traits
1/7/2019 The right - and wrong - way to use DNA tests
1/6/2019 Are breeding restrictions putting your breed at risk?
12/31/2018 More on "Simple strategies to reduce genetic disorders in dogs"
12/29/2018 Simple strategies to reduce genetic disorders in dogs
12/24/2018 Celebrating the preservation breeder!
11/27/2018 On preserving the purebred dog.
11/23/2018 Is the Ky allele in Wirehaired Pointing Griffons evidence of cross-breeding?
9/7/2018 Cool tricks with Kinship Coefficients, part 4: How closely related are the dogs in my breed?"
9/6/2018 Cool tricks with Kinship Coefficients, part 3: "How can I manage a disease without a DNA test?"
9/4/2018 Cool tricks with Kinship Coefficients, part 2: "Should I breed this dog?"
9/4/2018 Cool tricks with Kinship Coefficients, part 1: "Is this dog really an outcross?"
8/22/2018 The easy way to understand inheritance of recessive alleles
8/10/2018 The amazing secrets hiding in your pedigree database
7/21/2018 We can reduce the risk of hip dysplasia NOW!
7/12/2018 Is BetterBred better?
7/7/2018 Assessing genetic diversity and relatedness in dogs using DNA
6/30/2018 Using genomics to manage genetic disease. You don't need to find the genes
6/28/2018 How much does outcrossing improve genetic diversity?
6/26/2018 Are you improving genetic diversity, or just pushing the peas around?
6/25/2018 NEW: ICB Genetic Management Workshops
6/21/2018 A DNA Primer for Dog Breeders. Genetic Diversity: Inbreeding (Fis)
6/21/2018 A DNA Primer for Dog Breeders. Genetic Diversity: Inbreeding (ROH)
6/21/2018 A DNA Primer for Dog Breeders. Genetic Diversity: Heterozygosity
6/21/2018 A DNA Primer for Dog Breeders. ICB Breeder Tool Quick Start Guide
6/21/2018 A DNA Primer for Dog Breeders (You have your dog's DNA data. Now what?)
6/17/2018 No pedigree? No problem!
5/31/2018 A key innovation in dogs: diet
5/1/2018 The lesson(s) from SOD1and degenerative myelopathy
10/27/2017 Update on Newfoundlands
10/26/2017 Please don't ruin the Newfoundland
8/26/2017 The amazing dog nose: can you smell me now?
8/24/2017 The complexity of cancer
8/12/2017 Are preservation breeders preserving the Doberman? (No.)
8/5/2017 Hip laxity and the risk of degenerative joint disease
8/2/2017 Making better decisions about hip and elbow dysplasia: the era of genomics is here
4/29/2017 New insights into the development of dog breeds
4/27/2017 The genetic status of the Bernese Mountain Dog
4/3/2017 How to win The Health Test Game
3/12/2017 An update on the genetic status of the Doberman Pinscher
3/9/2017 Lessons from wolves
3/6/2017 Why "vulnerable breeds" are vulnerable
3/3/2017 Inbreeding and the immune system: unintended consequences
3/1/2017 The questions PUPscan won't answer. Part 2: The answers
2/28/2017 The questions PUPscan won't answer. Part 1
2/5/2017 Latest OFA statistics for hip dysplasia (Dec 2016)
2/2/2017 Why didn't Antarctic sled dogs have hip dysplasia?
1/23/2017 Your handy DNA testing crib sheet
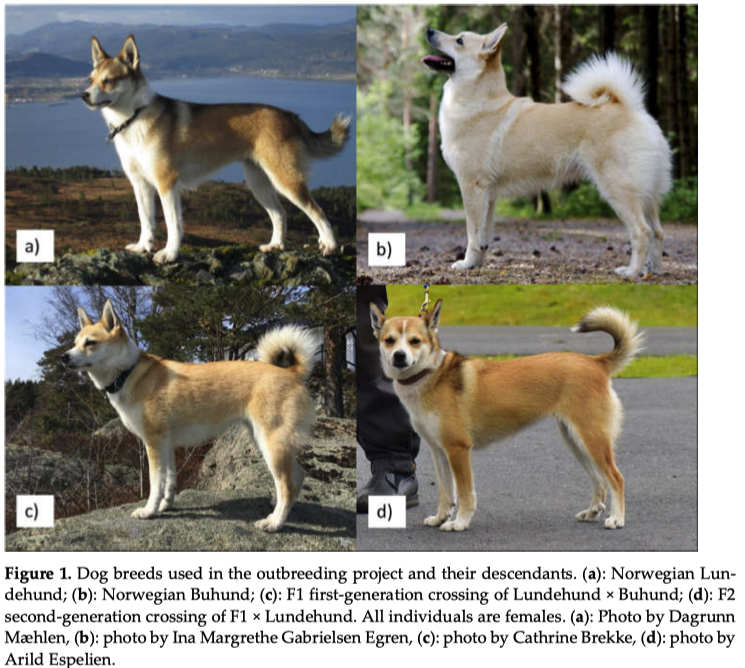
1/18/2017 Rescuing the Norwegian Lundehund: an update from Milo
1/3/2017 Comparing levels of inbreeding in dogs and horses
12/26/2016 Inbreeding of purebred dogs determined from DNA
12/15/2016 NEW: ICB Genetic Diversity Certification
12/9/2016 Why we need a more wholistic approach to managing canine genetic disorders
12/7/2016 A simple new tool for genetic disease management
12/4/2016 The ICB Breeder Tool: Overview
11/26/2016 Dog breeding in the era of genomic selection
11/23/2016 The new ICB Genomic Breeding Tool: the Genomic Relationship Coefficient
9/18/2016 How to develop effective strategies for the genetic management of your breed
9/2/2016 Preventing transmission of infectious disease at dog shows and sporting events
8/31/2016 Gone too soon? Enough already.
8/27/2016 Hip dysplaysia facts, fallacies, and fairy tales
8/16/2016 Why you should care about effective population size
8/14/2016 The world's oldest cancer...in dogs
8/7/2016 Introducing a new course: The Biology of Dogs
7/29/2016 Bulldog breeders: a call to action
7/23/2016 Try these breeding games!
7/4/2016 Genes and the amazing mind of the dog
7/2/2016 A game-changer for breeders: the ICB Breeder Tool
6/30/2016 Understanding the heritability of behavior in dogs
6/24/2016 Certificate of Completion: Genetics of Behavior & Performance course
6/5/2016 Are we watching the extinction of a breed? (part 2)
6/4/2016 Are we watching the extinction of a breed?
4/15/2016 A broader view of extinction risk of dog breeds in the UK
4/2/2016 A call for preservation breeding
3/29/2016 Twenty key elements of a successful breeding program
3/28/2016 Breeds with the BEST & WORST genetic diversity
3/25/2016 What are we going to do about Terriers?
3/20/2016 Evaluating the genetic status of a breed using both pedigrees and DNA
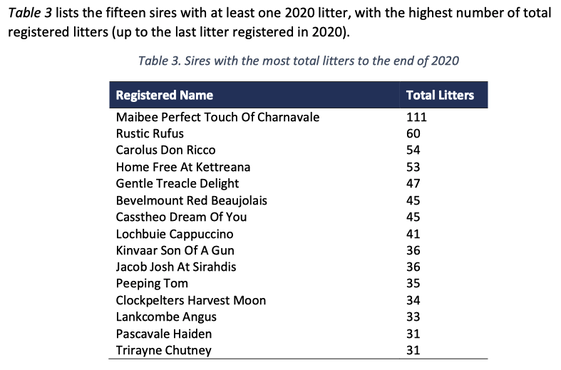
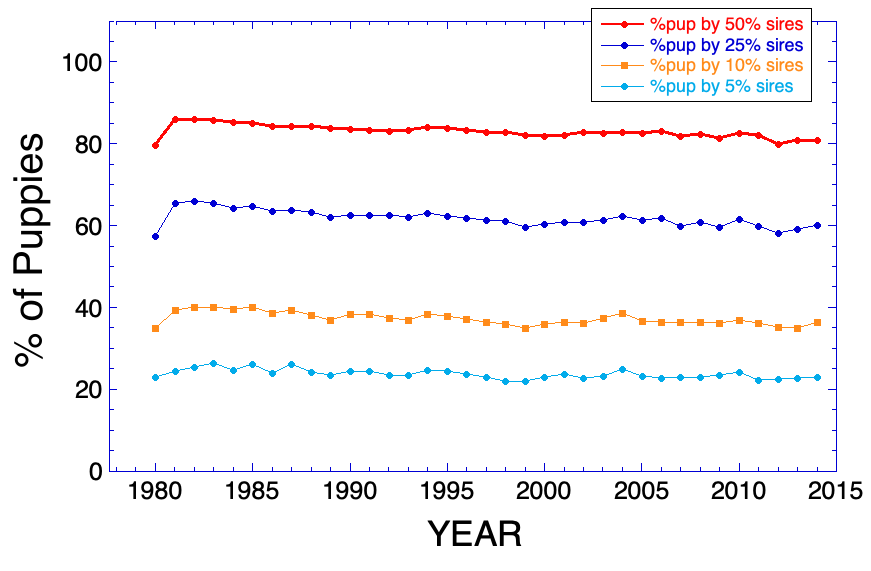
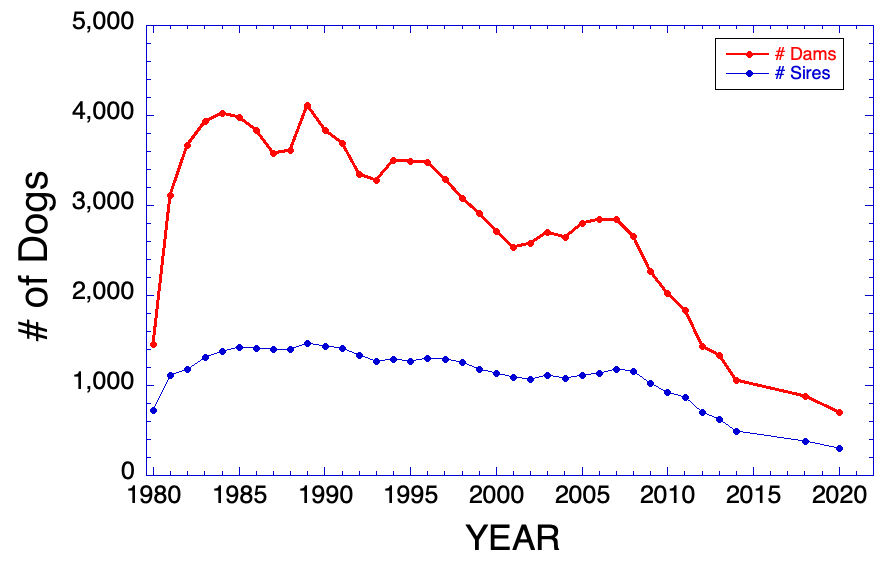
3/15/2016 Reprise: The Pox of Popular Sires
3/13/2016 That purebred vs mixed breed thing again
2/7/2016 Do you know what you don't know?
1/31/2016 Do you REALLY need to take a genetics course?
1/27/2016 Three key strategies to reduce genetic disorders in dogs
1/17/2016 Is it Nurture or Nature?
1/2/2016 Managing risk factors for hip dysplasia
12/23/2015 How do hips become dysplastic?
12/21/2015 Reliability of DNA tests for inherited diseases in dogs
12/16/2015 Virtual tours of the canine hip and pelvis
12/11/2015 The 10 most important things to know about canine hip dysplasia
11/4/2015 Coming soon: Course Certifications!
11/2/2015 Brachycephaly: it's more than just the pretty face
10/24/2015 The poop about dog diets
10/12/2015 Is (raw) diet the problem?
10/10/2015 Do dogs have more cancer than other mammals?
9/29/2015 Myths and mysteries about hip dysplasia
9/21/2015 Genetic status of purebred dogs in the UK
9/16/2015 Bigger puppies develop hip dysplasia
9/14/2015 The Mongolian Bankhar Dog Project
9/5/2015 Citizen Scientists: Let's do something about hip dysplasia!
9/1/2015 Major 2015 epilepsy consensus report
8/26/2015 Genetics, behavior, and puppy temperament testing
8/24/2015 The problem with the immune system: if you break it, it's yours
8/22/2015 Managing genetic disorders: "Just eliminate the bad gene"
8/9/2015 Is the dam more important than the sire?
7/8/2015 Decoding the genetics of behavior in dogs
6/23/2015 Looking for early pedigree data?
6/14/2015 For genetic improvement, it's the mix that matters
6/12/2015 The relationship between inbreeding and genetic disease
6/9/2015 Putting dogs to work for conservation
6/4/2015 COI FAQs: Understanding the Coefficient of Inbreeding
6/2/2015 Solving the problem of genetic disorders in dogs
5/14/2015 Visualizing inbreeding on the chromosome
4/30/2015 The trouble with Terriers
4/29/2015 Vulnerable breeds: how small is too small?
4/1/2015 A bright future for purebred dogs
3/29/2015 Health of purebred vs mixed breed dogs: the actual data
3/27/2015 Finding genes without DNA
3/26/2015 Tracing the paths of drifting genes
3/24/2015 If knowledge is power, know every puppy
3/19/2015 Lush on linebreeding
3/12/2015 Why all the fuss about inbreeding? (Or "Why are there so many genetic disorders in dogs?")
3/7/2015 What does "health tested" really mean?
1/9/2015 The history of purebred dogs in the UK
1/5/2015 Genetic test for renal dysplasia (Caution advised)
1/2/2015 A better way to pick 'em: using EBVs to reduce genetic disorders in dogs
1/1/2015 Estimating the breeding value of a dog
12/31/2014 Why do dogs get cancer?
12/28/2014 Cryptorchidism is complicated
12/26/2014 Silent secrets in old dog bones
12/22/2014 The myth of hybrid vigor in dogs...is a myth
12/17/2014 Hitting the bottle: the genetics of boom and bust
12/4/2014 More on tending the genetic pantry
12/1/2014 Using inbreeding to manage inbreeding
11/25/2014 Why dogs are sloppy drinkers (and cats aren't)
11/21/2014 The complexity of coat color
11/18/2014 Epilepsy incidence and mortality in 35 dog breeds
11/9/2014 Reducing genetic risk
11/7/2014 Take the breeder quiz!
11/6/2014 Dealing with those pesky mutations
10/31/2014 It's not always as simple as dominant and recessive
11/9/2014 The fiction of "knowing your lines"
10/24/2014 Is your breed drifting?
10/23/2014 Who's tending your genetic pantry?
10/15/2014 How breeding the best to the best can be worse
10/10/2014 When Should You Spay or Neuter Your Puppy?
10/3/2014 Genetic disorders in dogs: breaking the machinery of life
9/25/2014 Get Started Using Estimated Breeding Values (EBVs)
9/19/2014 The Costs and Benefits of Inbreeding
8/27/2014 A bit of sheepish fun
8/20/2014 How many generations of pedigree data should you use to estimate inbreeding?
7/23/2014 Me, jealous? Never! But my dog, on the other hand...
7/20/2014 Population Size & Inbreeding
7/19/2014 Avoiding inherited genetic diseases in dogs
6/18/2014 Wright’s Coefficient of Inbreeding
6/5/2014 Why DNA tests won't make dogs healthier
6/1/2014 Eliminating genetic disorders in dogs - too little, too late?
5/2/2014 Better hips and elbows? Maybe.
5/1/2014 Cancer Surprises
2/21/2014 Genetic Management of Dog Breed Populations
2/2/2014 What Does Population Genetics Have To Do With Breeding Dogs?
12/5/2013 The Pox of Popular Sires
10/5/2013 A bit more about Poodles
9/24/2013 An open letter to the Canadian Poodle clubs and others that love the breed
7/23/2013 Why do dogs have so many genetic disorders?
7/19/2013 Primary lens luxation is WIDESPREAD among dog breeds - are you testing?
7/18/2013 Inherited myopathy in Labradors is found worldwide - the legacy of a popular sire
7/2/2013 Finally, a scientific journal about Dogs!
3/9/2013 How molecular genetics will change dog breeding
7/6/2012 Locating the genes for hip dysplasia in dogs (Psssst! Look in the kibble bag)
4/19/2012 Population genetics suggests dire straits for Tollers and Heelers