| The number of breeding animals in a population can have profound effects on population genetics. Let's say we have a herd of 10,000 antelope. If a few of those animals fail to reproduce successfully, perhaps because they are too old or their offspring were eaten by wolves, the consequences to the genetics of the population as a whole will be negligible because there is such a large number of animals. The loss of an animal or two here or there from a population of many thousands is not going to reduce the number of potential breeding partners and is unlikely to result in the loss from the population of the very last copy of some particular allele. But what if the size of the herd is only 10? Now, the loss of a single animal reduces the population size by 10%, which means there are fewer breeding options for the next generation, and over time this will increase the rate of inbreeding. Also, the animal that is lost could have been carrying the only copy of a rare allele and it is lost from the population forever. |
Genetic drift is the random variation in allele frequencies from one generation to the next that occurs in populations. Genetic drift is what causes groups of animals separated from the main population to drift apart genetically over time. Drift can result in the loss of alleles just by chance, or it can cause "fixation" of an allele, in which every member of the population is homozygous.
Rate (inbreeding or genetic drift) = 1/2Ne
where Ne is "effective population size", which we will define shortly.
As an example, if we have a population of 10 breeding animals, the rate of inbreeding will be 1/(2 x 10) = 1/20 = 0.05. As a percentage, that would be a 5% increase in homozygosity per generation, which is very high. What if the population size is 100? The rate of inbreeding would be 1/(2 x 100) = 0.005, or 0.5% increase in homozygosity per generation. For genetic diversity, the calculations are the same, and the value is the amount of genetic diversity (as a fraction or a percentage) lost per generation.
So, the rates of inbreeding and of genetic drift per generation will increase as Ne gets smaller.
EFFECTIVE POPULATION SIZE
If you are managing groups of animals, you might want to understand how the size of a particular population will affect genetics. The easiest way to do this is to pretend you have an "ideal" population that meets certain criteria:
a) The number of males and females is equal and all can reproduce;
b) All animals are equally likely to produce offspring, and the number each produces varies no more than expected by chance;
c) Mating is random (remember Hardy-Weinberg?);
d) The number of breeding animals is the same from one generation to the next (i.e., the population size is constant) and there is no immigration, emigration, mutation, or selection.
For this ideal population, we can determine something called "effective population size", Ne.
In an "ideal" population (one that meets the criteria listed above), the census population size and the effective population size are equal. But most real populations violate at least some of the criteria for an ideal population (e.g., sex ratios are not equal, breeding is not random, etc), and for them the effective population size is less than the census size.
If real populations are rarely ideal, what's the point of having something called effective population size? The genetics of ideal populations are predictable. If we could describe our herd with a census size of 10,000 antelope in terms of effective population size, we could make predictions about how it would behave genetically. How can we get from census size, Nc, to Ne?
There are several ways to estimate Ne for a population, depending upon the data available and the type of estimation you're trying to make.
First, let's produce a formal definition of Ne:
Effective population size is the size of an "ideal" population of animals that would have the same rate of inbreeding or decrease in genetic diversity due to genetic drift as the real population of interest.
Effect of sex ratio on Ne
There are several different ways to compute Ne. We will look only at one here that is easy for breeders to use.
One of the things that can influence the effective population size is the sex ratio of the breeding animals.
We can estimate Ne using information from a population census or pedigree database about the numbers of males (Nm) and females (Nf) that produce offspring in a generation.
Ne = 4 x (Nm x Nf) / (Nm + Nf)
The denominator (bottom) of this equation is the number of males plus the number of females (Nm + Nf), which is simply the total number of breeding animals. The numerator (top) of the equation is four times the product of Nm and Nf (4 x (Nm x Nf)).
A simple example
Let's say we have a population of 10 animals, 5 males and 5 females. The effective population size Ne would be
Ne = 4 x (5 x 5) / (5 + 5)
Ne = 4 x 25/10 = 10
See that? When we had equal numbers of males and females, Ne is the same as the census size. Try working a few of these with other population sizes, but always with equal numbers of males and females.
As you know, the number of breeding males and females in purebred dog populations are rarely equal, with the number of females usually being greater because males can sire multiple litters. What happens to the effective population size when the sex ratio of breeding animals is not equal?
An extreme example
Let's just cut to the chase and create a crazy population of animals, with 1000 females and just 1 male. For this, the math will be:
Ne = 4 x (1000 x 1) / (1000 + 1) = 4 x 1000/1001 = 3.996, or approximately = 4.
Wow. We have 1,001 animals, but the effective population size is FOUR.
We had a herd of 1,000 animals, and because we used only one male for breeding, our population is going to behave genetically AS IF it is a population of only 4 animals, two males and two females.
What if there are 1,000 males and only one female? Apart from the awkward logistics, the result in terms of genetics will be the same. The Ne will again be approximately 4.
As you can see, an imbalance in the number of breeding males and females will affect the estimate of Ne. How big this effect is will depend on the size of the population and the sex ratio.
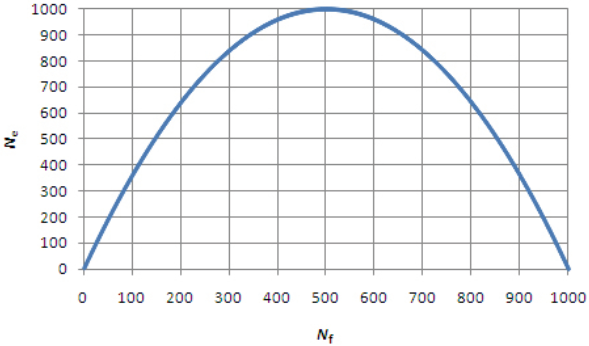
Let's look at this graph, which is for a population with a total of 1,000 animals. The number of females (Nf) is on the x-axis, and the number of males is not displayed but inferred by subtraction (Nm = 1000 - Nf). When the number of males and females is equal (i.e., Nf = Nm = 500), the effective population size (Ne) equals 1,000; i.e., Ne is the same as the census population size.
When the number of females (Nf) is 800 (so the number of males (Nm) is 200 by difference), the resulting effective population size from the graph is about 650. If the number of breeding males is reduced to 100, Nf will be 900 and Ne will be about 350. The curve is symmetrical; you can run similar examples with small Nf and large Nm, and the resulting Ne will be the same.
What you can see here is that when the sex ratio of breeding animals is not 1:1, the Ne is reduced, meaning that rates of inbreeding and genetic drift will be similar to the ones you would expect to see in a population of smaller size. Furthermore, the more extreme the ratio, the greater the effect on the genetics of the population.
I'm sure you can see the problem here. Inbreeding and loss of genetic diversity result in an increase in inbreeding depression and the expression of recessive mutations. Litter sizes get smaller, puppy mortality goes up, lifespan is shortened, and unhealthy animals are removed from the breeding stock. Reducing the number of breeding animals will cause the rates of inbreeding and genetic drift to increase, which will increase inbreeding depression and the expression of genetic diseases, which will reduce the number of breeding animals...and you can see where this is going. This phenomenon is called the "extinction vortex".
You can also see how allowing only a few of the males to breed (or a popular sire) will affect the gene pool. While breeders think they are improving genetic health by using only the very best dogs for studs, the result is that inbreeding and loss of genetic diversity will both increase, and you will end up in the extinction vortex.
There are consequences here for spay/neuter policies and breeding contracts. Removal of animals from the breeding population makes it more difficult for breeders to manage the rate of inbreeding, and higher levels of genetic drift cause genetic instability, with larger fluctuations in allele frequencies and greater risk of extinction.
This raises an important consideration when using information from DNA trait and mutation tests and genotyping (e.g., MyDogDNA, UC Davis diversity test) to make breeding decisions. Breeders are able to improve breeding decisions through careful mate selection using DNA information, but if the sex ratio of breeding animals is strongly unbalanced or the size of the breeding population is too small, inbreeding and loss of genetic diversity will be working in the opposite direction, compromising the health of the gene pool. Genotyping can help improve the quality of the next litter, but reducing the incidence of genetic disorders in a breed will require appropriate genetic management of the entire population using breeding strategies that will limit the rate of inbreeding and loss of genetic diversity. In fact, if breeders are not keeping an eye on the larger picture, these tests might cause breeders to be even more selective in their breeding choices, with deleterious long term consequnces. In fact, this is what happened when genomic testing was first introduced to livestock breeding. Selecting only the best animals for breeding based on DNA increased the rate of inbreeding and loss of genetic diversity, producing short term gain but damaging the gene pool. There is no silver bullet that will cure all problems. Breed-level genetic management is essential to maintain the health of the gene pool over the long term.
Breeders should know the effective population size of their breed, and they should understand the things they can do to increase it. Larger Ne will improve genetic stability and the health of the gene pool; smaller Ne will result in unpredictable variation in allele frequencies, loss or fixation of some alleles, and an increase the risk of extinction. You can see how I was able to use information from the UK Kennel Club stud books to estimate effective population sizes of dog breeds in the UK and particularly of terriers, and from this produce lists of breeds at risk of extinction.
Although Ne is not especially hard to compute with the formula that was used here, there is an online calculator that will allow you to experiment with sex ratio and population size: Effective Population Size Calculator.
ICB's online courses
***************************************
Visit our Facebook Groups
ICB Institute of Canine Biology
...the latest canine news and research
ICB Breeding for the Future
...the science of dog breeding